Generating new Protein Sequences with Language Models
"Uncovering the Science behind Life with AI" series - Part 1
This is the first post in the series “Uncovering the science behind life with AI,” in which we will present applications of artificial intelligence in the field of life sciences. We will begin by describing protein language models, and will present in more detail a recent model that is particularly interesting because the protein sequences it generated were also synthesized and tested in the laboratory.
Proteins are all around us. Not only do they build up the cells and tissues of all living organisms, but we often use isolated proteins in our everyday lives. Among other things, proteins are found in detergents, are used to process food, and are also used as medicines or diagnostic tools (e.g., in rapid antigen tests). However, it happens that for some applications, we need a protein with certain properties that no known protein fulfills. For example, we need an enzyme that can effectively catalyze a chemical reaction at lower temperatures or one that is capable of degrading a synthetic organic molecule. Or we would like an antibody that binds to a pathogen or a molecule associated with a disease to develop a diagnostic tool or even a therapy. However, designing new proteins with the desired properties is a difficult task. Traditionally, this has been accomplished by iteratively altering individual amino acids in the sequence of a natural protein or by generating new sequences unrelated to those found in nature based on the physical principles of intramolecular and intermolecular interactions. However, the latter approach is currently applicable to short proteins only due to the high complexity of such models. Recently, deep language models have been developed as an alternative for generating new proteins. But before we take a closer look at these so-called protein language models, we should first clarify what proteins actually are and why language models are suitable for proteins.
Proteins as Biological Macromolecules
Proteins are biological macromolecules with multiple functions in cells and the body. Some proteins, called enzymes, carry out chemical reactions, such as the enzymes in saliva, the stomach, and the intestines that break down the food we eat. Then there are many proteins that provide structure and support, e.g., the robust protein keratin, which is found in hair, nails, and the outer layer of the skin. Other roles of proteins include storage and transport of compounds (remember hemoglobin, the protein that carries oxygen to cells), signal transduction to coordinate biological processes, and immune response (e.g., antibodies that recognize pathogens).
Proteins are made up of building blocks called amino acids. Almost all the proteins we know of are made up of exactly twenty different amino acids. Each amino acid has its own physical and chemical properties (e.g., whether it is polar or nonpolar), and based on these properties, we can divide them into groups of amino acids with similar properties.
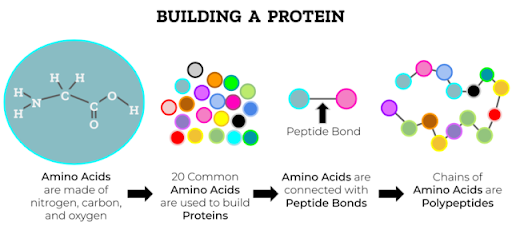
Amino acids in a protein are connected linearly and form chains. The order in which the amino acids appear in the protein chain is called “sequence” of the protein and can be represented as a single long word, a string of twenty different letters, with each letter representing an amino acid. The protein sequence determines all the properties of the protein: What 3D shape the chain of amino acids folds into, the catalytic properties of enzymes, how strongly it will bind to other molecules, etc. In different organisms, we can find similar (but not identical!) proteins that have similar sequences and 3D structures and usually perform similar functions. Such proteins emerged from a common evolutionary origin, but then changed slightly due to mutations in each individual organism. We say that these proteins belong to the same protein family.
As mentioned earlier, proteins of the same protein family have similar sequences. In each protein, there are some amino acids in specific locations that are most crucial for the final function of the protein – if you were to replace that amino acid with another amino acid with completely different properties, it would (most likely) result in a loss or alteration of function. Therefore, these crucial amino acids are usually well conserved in all members of the protein family or are replaced by similar amino acids in some proteins of the family. In a typical protein, there are usually some regions with well conserved amino acid sequences, while in the rest of the protein sequence, the composition and order of amino acids vary more.
Proteins as Strings of Amino Acids
In fact, we can see that there are some similarities between natural language and protein sequences: Both can be represented as strings of letters, and both are composed of reusable modular elements that can be assembled into larger units according to certain rules. In the case of proteins, the letters denote amino acids, while the modular elements are the well-preserved amino acid sequences, analogous to words, phrases, and sentences in natural language. Another key feature that proteins and natural language have in common is the completeness of information – as mentioned earlier, the amino acid sequence determines all the properties of the protein. Because of the similarities in form and content between protein sequences and natural language, natural language processing (NLP) methods can be successfully applied to protein sequences to study local and global properties [2].
However, there are also some important differences between natural language and protein sequences. Most human languages have punctuation that clearly separates the individual structures, whereas proteins generally lack such clearly separable structures, and the amino acid sequences of the protein’s functional units also often overlap. Proteins also lack a clear vocabulary. In addition, proteins exhibit high variability in length: The shortest proteins consist of a few dozen amino acids, while the length of the longest proteins is in the tens of thousands of amino acids. Last but not least, in proteins, there are much more distant interactions between amino acids that are far apart in the linear sequence but spatially close in the 3D structure of the folded protein [2].
Protein Language Models
Deep learning models based on natural language processing methods, in particular attention-based models that are trained on large collections of protein sequences instead of text, are called protein language models. During training, they capture local and long-range relationships between amino acids in a protein sequence. Finally, protein language models learn the probability of an amino acid or amino acid sequence occurring in a protein and are able to generate new protein sequences.
ProGen
In the following part of the blog post, we will introduce in more detail a recent protein language model called ProGen [3], which has also been experimentally validated, i.e., actual proteins were synthesized in the laboratory from some of the generated protein sequences, and the synthesized proteins also functioned in the expected manner.
ProGen is a transformer-based conditional language model trained unsupervised on 280 million protein sequences from publicly available datasets prepended with control tags. The control tags are metadata associated with the protein sequences from the datasets, such as which organism the protein is from, which protein family it belongs to, the function of the protein, the biochemical process or cellular component it is associated with, etc. These control tags are then used for conditional generation of protein sequences, i.e., ProGen generates protein sequences with varying degrees of similarity to natural proteins based on control tags provided as input (prompts) to the language model.
ProGen’s transformer architecture consists of 36 layers with 8 self-attention heads per layer and a total of 1.2 billion trainable neural network parameters. The self-attention mechanism in each layer derives pairwise interactions among all positions in its input sequence. By stacking multiple self-attention layers, the model can learn multiple amino acid interactions, including long-range interactions.
After training the model, ProGen can be fine-tuned (i.e., further trained) using sequences from protein families of interest so that ProGen can capture local sequence specificities for these protein families and further improve the quality of the sequences generated by the model. Fine-tuning involves small, low-cost gradient updates to the parameters of the trained model.
In the research article published in Nature Biotechnology that featured ProGen [3], the authors fine-tuned their model to lysozymes (enzymes found in tears and saliva, for example, that break down bacterial cells to defend against microorganisms) and then generated 1 million sequences using the lysozyme protein family identifier as a prompt. From the 1 million sequences generated, they selected 100 sequences with varying degrees of sequence similarity (40–90%) with the natural lysozyme based on a combination of an adversarial discriminator and a generative model with log-likelihood assessment. They trained the adversarial discriminator to distinguish between natural and ProGen-generated lysozymes, and a higher discriminator score was given to the generated protein sequence, which should be functionally closer to the natural lysozymes but does not necessarily have very high sequence similarity to the natural lysozymes.
The scientists synthesized the selected 100 ProGen-generated proteins in the laboratory, along with 100 natural lysozymes that served as controls. They then tested both the ProGen-generated and natural lysozymes for their ability to break down bacterial cells using a standard laboratory assay. The generated and natural proteins performed equally well.
This proof-of-concept study demonstrates that a deep language model trained solely on an extensive collection of protein sequences is capable of generating new protein sequences for functional proteins with the specified properties without additional information about the structure or evolutionary relationships among proteins. In general, protein language models are emerging as a good alternative approach for designing new proteins and could be used in the future to create new proteins that, as mentioned at the beginning of this post, are needed in various applications, ranging from basic biochemical research to developing new therapies or more environmentally-friendly solutions.
Angelika Vižintin
Angelika Vižintin received her PhD in Biosciences at the University of Ljubljana, Slovenia. In her PhD thesis, she studied the effects of short high-voltage electric pulses on animal cell lines and their potential application, for example, for electrochemotherapy, a local cancer treatment that combines membrane-permeabilizing electric pulses and chemotherapeutic drugs. She is currently taking courses in machine learning, mathematical foundations of artificial intelligence, computer science, etc. in the Artificial Intelligence programme at Johannes Kepler University in Linz, Austria, with the goal of learning how to use artificial intelligence tools in biological/biomedical research. She also produces science-related programs for the Slovenian Radio Študent, one of the oldest and strongest independent radio stations in Europe. Among the topics she reports on most frequently are the biology of the female body and women’s diseases, biotechnology, genetically modified organisms, molecular biology, and artificial intelligence in the life sciences. In her free time, she enjoys listening to music and traveling.
References
[2] Ofer D, Brandes N, Linial M.
The language of proteins: NLP, machine learning & protein sequences.
Comput Struct Biotechnol J. 2021 doi: 10.1016/j.csbj.2021.03.022.
[3] Madani A, Krause B, Greene ER, Subramanian S, Mohr BP, Holton JM, Olmos JL Jr, Xiong C, Sun ZZ, Socher R, Fraser JS, Naik N.
Large language models generate functional protein sequences across diverse families.
Nat Biotechnol. 2023 Jan 26. doi: 10.1038/s41587-022-01618-2.

